Portable Molekel 5.3
Portable Molekel 5.3 :: 48,4 MB
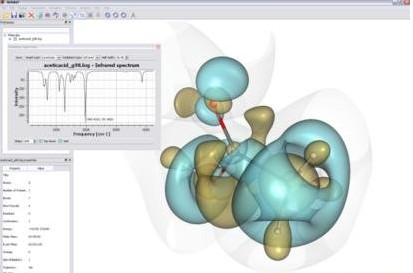
MOLEKEL is an interactive molecular graphics program to visualize molecular and electronic structure data from a number of electronic structure program outputs (Gaussian, Gamess, ADF…) as well as from XYZ and PDB files. As one of the major new features MOLEKEL can now also read Gaussian cube files. MOLEKEL is able to render the molecular structure in different styles, display orbitals, electron densities, map and color code properties on any surface, display and animate vibrational normal modes, animate a sequence of coordinates and produces great graphics.
Some of the features available in the new version are:
* Multiplatform: Mac OS X, Windows, Linux
* Completely redesigned UI
* Different methods to speed-up rendering of molecules with support for billboards and view-dependent level of detail techniques
* Programmable shaders; standard shaders to enhance rendering quality, outline contours and perform sketch-like renderings are provided
* Visualization of residues (ribbon or schematic)
* Complete control over the generation of molecular surfaces (bounding box and resolution)
* Visualization of the following surfaces:
o Orbitals
o Iso-surface from density matrix
o Iso-surface from Gaussian cube grid data
o SAS
o SES
o Van der Waals
* Animation of molecular surfaces
* Animation of vibrational modes
* Export to PostScript and PDF
* Export animation
* Plane widget to visualize a scalar field: the plane can be freely moved in 3d space and the points on the plane surface will be colored according to the value of the scalar field: a cursor can be moved on the plane surface to show the exact value of the field at a specific point in space.
* Fully Doxygen-commented source code.
Download: (Size: 48,4 MB)
RapidShare
Portable Molekel 5.3 için yorumlar kapalı

